 
As a result, both the top and bottom DNA strands are cut and only held together by the few hydrogen bonds between the 5'-AATT-3' on both strands. In the example of EcoRI, this enzyme cuts the DNA between the "G" and the "A" on the 5' side. The rest of the restriction enzyme name comes from the strain of the organism (in this case the R strain) and the order in which the enzyme was discovered (in this case it is the first restriction enzyme discovered from the R strain of E. The " Eco" part of the enzyme name comes from the fact that this enzyme originates from Escherichia coli ( E. This example is a recognition sequence from the restriction enzyme known as EcoRI. Notice that the complementary DNA strands above, if reading from the 5' end, have the same sequence: 5'-GAATTC-3'. Ordinary words that are palindromic include "mom," "dad," "wow," and "racecar." With DNA, a palindrome is based on reading one DNA strand 5' to 3' and comparing it with its complement DNA strand as read 5' to 3'. Typically, recognition sites are palindromic, that is they read the same backwards and forwards. Recognition sites are different for each restriction enzyme. These enzymes cut DNA at a characteristic recognition site. Restriction enzymes are found in some bacteria and have been isolated to use for a variety of biotechnologies such as DNA fingerprinting. For paternity testing, half of the fingerprint will originate from the biological mother and half of the fingerprint will originate from the biological father. If fingerprints match, it likely means that the DNA originated from the same organism. The result is a pattern of bands that can be compared with other patterns from known samples. The DNA fragments produced by the restriction enzyme are separated by size using an approach called gel electrophoresis ( see the Gel Electrophoresis section below). The restriction enzyme will cut the DNA in a pattern that will differ from DNA from other sources, unless the identify of the DNA is the same (matching known and unknown samples enables identification). DNA that has been removed from cells and other cell components) is mixed with a restriction enzyme to create a fingerprint. 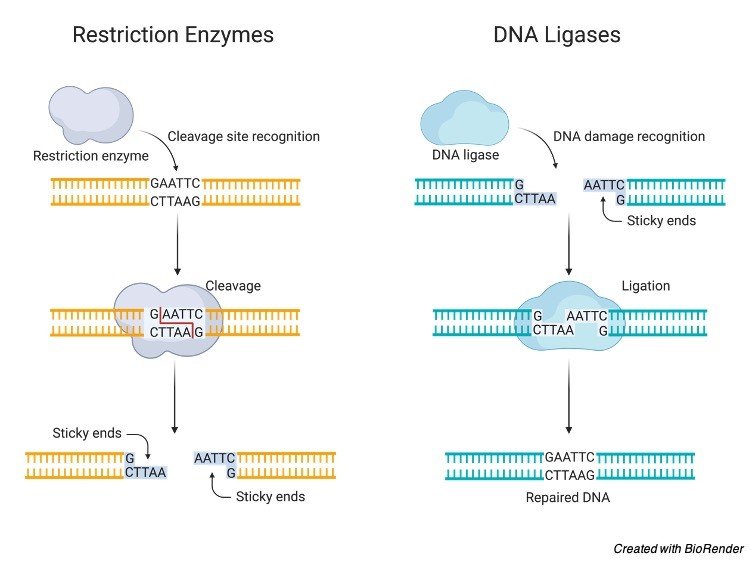
If the samples match, it enables identification. \)ĭNA fingerprints are created by first isolating DNA from an unknown sample to be identified and compared with known samples.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
